-
Early Research
-
Dissertation Research
-
Post-doctoral Research
-
Future Research
<
>
Early Research Experience
|
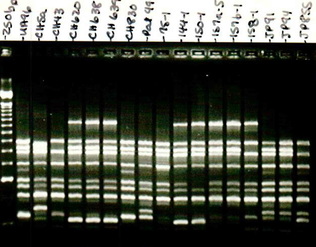
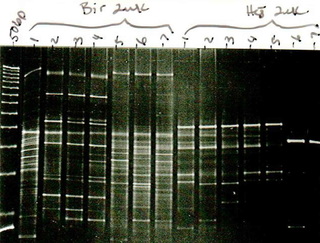
Before entering the PhD degree program, I held three positions where I had the opportunity to investigate diverse areas. My first position was a research technician in the laboratory of Dr. Page Caufield under the supervision of Dr. Yihong Li. My research projects in Dr. Caufield’s laboratory included AP-PCR analysis of Streptococcus mutans, denaturing and temperature gradient gel electrophoresis for community profiling of bacteria from oral, gut, and surgical sites. I also learned basic laboratory skills for PCR, gene cloning, whole chromosomal DNA fingerprinting, and enzyme digests.
|
|
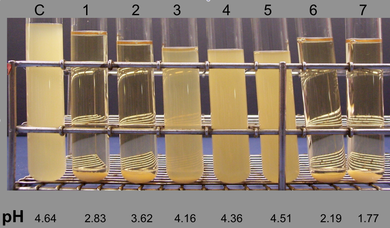
My second position was as a Program Coordinator managing a non-Profit UAB Service Center University Sterilization Assurance (USA), which later merged into a Quality Assurance Officer for the School of Dentistry. While this position was not specifically a research position, I was offered several opportunities to work on various dental and environmental microbiology projects with dental school residents and as well as independently under the supervision of Dr. John Ruby. My research projects included affect of pH on S. mutans agglutination, effect of xylitol on S. mutans acid production from glucose and pH studies of candies & beverages. I also performed routine testing for heterotrophic bacteria in dental unit waterlines, specialized Legionella testing of hospital water sources, biological monitoring of autoclaves, and contamination of soap dispensers. Most notably under Dr. Ruby’s supervision I was offered my first opportunity to work with a project using repetitive extragenic palindromic PCR (rep-PCR).
|
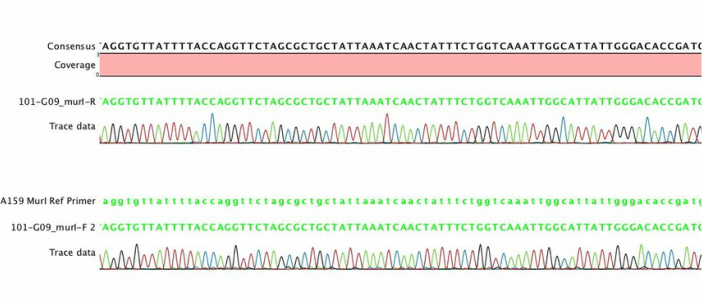
My third research position was as a research assistant under the supervision of Dr. John Ruby and Dr. Noel Childers. In this capacity I worked to isolate and genotype S. mutans in a large-scale (≈ 32,000 isolates) longitudinal epidemiological study. While my contributions to research up to this point were largely technical in nature, this period allowed me to develop my manuscript writing skills by contributing as a co-author for studies related to the early genotypic study of S. mutans using a new molecular based typing approach, repetitive extragenic palindromic PCR (rep-PCR). Our laboratory pioneered the use of this technology for the epidemiological study of S. mutans which offers a cost and labor effective alternative to pulse-field gel electrophoresis. In this position I completed my Masters of Science in 2010 working using multilocus sequence typing (MLST) to validate rep-PCR genotypes. This laboratory experience expanded my knowledge of sequence data processing (pictured in the header), real-time PCR, quantitative PCR, and clinical research data.
Dissertation Research
Upon entering the PhD program, I continued to work on the S. mutans epidemiological study in the laboratory of Dr. Noel Childers. However my focus shifted from evaluating these samples with rep-PCR to analysis with multilocus sequence typing (MLST). I also evaluated numerous selective media for the isolation of S. mutans and their effects on the genotype selection of the larger epidemiological study.
|
My doctoral dissertation investigated the genetic diversity and transmission of S. mutans using molecular analysis with rep-PCR and MLST for effective large-scale epidemiological study. Rep-PCR is a rapid, cost effective genotyping tool while MLST is a molecular based typing method with high discriminatory power. Together the two methods allow for rapid screening of the genetic diversity and transmission of S. mutans, which will be useful for epidemiological study of dental caries. Through this research, I expanded my research technical skills for genotyping including advanced sequence analysis, bioinformatics, database utilization and approaches for comparisons of genotyping methods.
|
The first two publications from my dissertation focused on addressing major concerns affecting genotyping with rep-PCR and MLST: choice of selective media and choice of MLST typing scheme, The first of these works confirmed that the selective Gold’s media (MSB) was as effective as other media in supporting mutans streptococci growth and that selection of media did not impact the growth of specific genotypes. The second paper compared two currently available MLST typing schemes for S. mutans and found these schemes to be comparable, therefore the scheme described by Nakano et al. (2007) was selected for all later MLST analysis in our laboratory.
The third publication from my dissertation investigated the clonality of randomly selected S. mutans isolates to confirm the clonality reported in my master’s work was not a result of sampling bias. This study confirmed a high degree of clonality and contrary to other published studies suggested that S. mutans can be geographically or ethnically distributed illuminating the importance of conducting localized studies as compared to global studies. Furthermore, this study reported the first account of S. mutans serotype k in a United States population opening a new and possibly clinically significant alternate avenue for interdisciplinary research for these S. mutans isolates, because serotype k has been linked to cardiovascular disease, hemorrhagic stroke, and inflammatory conditions.
The fourth publication focused on using multilocus sequence typing (MLST) to investigate the genetic diversity of S. mutans and its transmission among children and their household family members. The published work was unique in both its scale and scope, with over 13,000 S. mutans analyzed. The analysis was performed using a new perspective reporting 63% children share genotypes with their mothers, but 72% of children had at least 1 rep-PCR genotype not share with any household members. Children with multiple rep-PCR genotypes were found to be 2.3 times more likely to have dental caries.
The fifth publication confirmed rep-PCR as a good pre-screen of isolates for downstream applications and that rep-PCR, together with MLST, can be used to effectively track transmission sources for some S. mutans genotypes in epidemiological studies.
The third publication from my dissertation investigated the clonality of randomly selected S. mutans isolates to confirm the clonality reported in my master’s work was not a result of sampling bias. This study confirmed a high degree of clonality and contrary to other published studies suggested that S. mutans can be geographically or ethnically distributed illuminating the importance of conducting localized studies as compared to global studies. Furthermore, this study reported the first account of S. mutans serotype k in a United States population opening a new and possibly clinically significant alternate avenue for interdisciplinary research for these S. mutans isolates, because serotype k has been linked to cardiovascular disease, hemorrhagic stroke, and inflammatory conditions.
The fourth publication focused on using multilocus sequence typing (MLST) to investigate the genetic diversity of S. mutans and its transmission among children and their household family members. The published work was unique in both its scale and scope, with over 13,000 S. mutans analyzed. The analysis was performed using a new perspective reporting 63% children share genotypes with their mothers, but 72% of children had at least 1 rep-PCR genotype not share with any household members. Children with multiple rep-PCR genotypes were found to be 2.3 times more likely to have dental caries.
The fifth publication confirmed rep-PCR as a good pre-screen of isolates for downstream applications and that rep-PCR, together with MLST, can be used to effectively track transmission sources for some S. mutans genotypes in epidemiological studies.
|
I was fortunate to have the opportunity to work on a number of side projects independent of, as well as related to, my research during my PhD studies. One study investigated the antimicrobial effects of a purple violet pigment isolated from an Antarctic bacterium Janthinobacterium sp. Ant5-2 on S. mutans.
|
In another side study, we report an increased prevalence of S. mutans serotype k in our study population using three different sample types (plaque, saliva, and individual isolates) for analysis. This study is the first report of S. mutans serotype k in a US population and will improve sensitivity of published methods for serotyping by PCR.
Post-doctoral Research
Initially, I worked to collect preliminary data for my post doctoral research in the laboratory of Dr. Hui Wu in the Department of Pediatric Dentistry at the University of Alabama at Birmingham School of Dentistry. In 2020, we moved to Oregon Health & Science University in Portland, Oregon where I joined the Department of Integrative Biosciences in the School of Dentistry. Projects include strain-to-strain interactions of S. mutans and dental caries, impact of host immunity in dental caries, and relationship of small molecules in caries virulence. During my postdoctoral training I worked to acquire technical expertise in using oral microbiome analysis, whole genome sequencing of clinical S. mutans, biofilm analysis, gene mutagenesis, transcriptomics, metabolomics, and animal models (Drosophila and Rat Caries Models) while investigating how novel biosynthetic gene clusters or the presence of multiple S. mutans genotypes impact early childhood caries and overall human health. I am also added an immunological component to my training by investigating the relationship between oral bacteria (microbiome) and salivary IgA. During this training I worked to expand my grant writing skills leading to a successfully funded K-99/R00 application.
Comparative Genomics of Clinical S. mutans
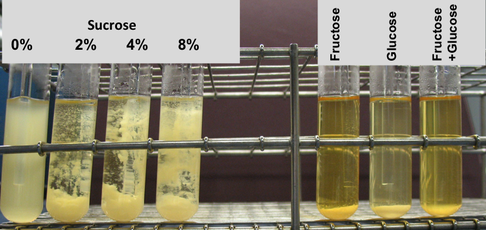
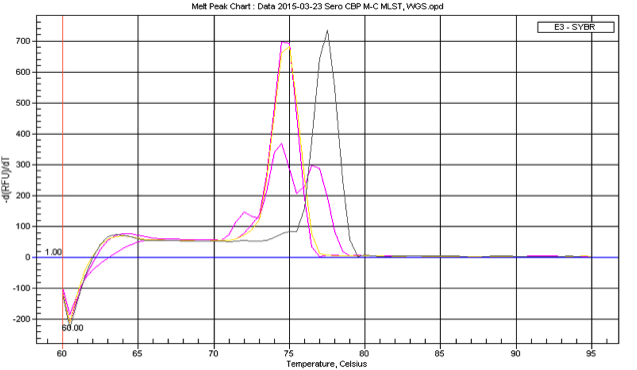
Comparative genomic analysis of whole genome sequences (WGS) for the 34 representative S. mutans rep-PCR genotypes reported in my dissertation was published in JDR in 2020. In combination with phylogenetic and phenotypic analysis (biomass, pH, bacteriocin production), WGS will allow us to characterize the commonality and diversity of virulence traits such as variability in the TnSmu2 operon (related to oxygen stress), diversity of mutacin profiles (related to survival against commensals), and variation in S. mutans virulence factors (e.g., ability to form biofilms, decrease biofilm pH).
Intra-species impact on caries virulence
My dissertation work indicated that children with multiple S. mutans genotypes were 2.3 times more like to have dental caries and other studies support this finding. It remains to be determined if this relationship causal or only an association. One of my current projects is investigating if the presence of multiple S. mutans genotypes have the ability to cause dental caries, or lead to more severe caries, using in-vitro and in-vivo models.
Host Immunity and Dental Caries
Although S. mutans is the primary bacterium most commonly associated with initiation of dental caries in children, other bacterial species can play an important role in dental caries disease progression. In our study population, we are interested in investigating the relationship of host immune response and oral microbiome.
Future Research
Novel small molecules of the Oral Cavity
Epidemiology of Childhood Dental Caries using Multi-omics.
The field of cariology has evolved from a single bacterial strain approach to polymicrobial models. However, emerging research is demonstrating that in order to advance even further, a systems biology approach for the epidemiological study of dental caries is needed. Using metagenomics, transcriptomics, and metabolomics, we can investigate the pathogenesis of dental caries across the spectrum. Bioinformatics data analysis pipelines are improving such that high-throughput meta-omics studies are becoming more manageable. Ultimately, I would like to pilot this study with a population, such as the Uniontown Study previously performed at the University of Alabama at Birmingham, then use this to apply for NIDCR R01 funding to expand this to other high-risk populations such as Native Americans and Hispanics. Our previous work has highlighted the importance of working within localized populations to attain achievable statistical power while reducing confounding variables. This longitudinal study will require a complex coordinated effort of many subject matter experts form the fields bioinformatics, pediatric dentistry, molecular biology, biochemistry, microbiology and community engagement. My previous experience in large scale epidemiological studies and meta genomic approaches as well as my project management skills will facility this successful multi-center collaboration.
Streptococcus mutans serotype k Prevalence
In my dissertation the first known S. mutans serotype k strains in the United States were reported. This opens a new avenue of research opportunities. We published a study to determine the overall prevalence of S. mutans serotype k in children in our study population using whole saliva, plaque samples, as well as individual bacterial isolates. We reported that testing multiple sample types will results in an even higher prevalence of serotype k than previously reported. Future studies are planned to investigate other populations to determine if the prevalence of serotype k observed varies between geographical or ethnic groups. Multidisciplinary teams to evaluate associations between serotype k, collagen binding proteins Cbm and Cnm, and systemic health will require successful collaboration with cardiovascular and neurology Departments of Medicine. An in-vivo model could validate other ongoing studies or demonstrate a direct causal link of serotype k to systemic health issues (cardiovascular disease, infective endocarditis, hemorrhagic stroke and inflammatory conditions).